The Global Garden project: Imagining plant science.
Nicholas M. Lee, Hannah E. Hodgson, Chris Hann, Mike O’Driscoll, Samantha Stebbings, Collette Matthewman, Miriam Kent, Jenni Rant and Anne Osbourn
Plants, People, Planet (2020) 2: 602-613
The Global Garden project: Imagining plant science.
Nicholas M. Lee, Hannah E. Hodgson, Chris Hann, Mike O’Driscoll, Samantha Stebbings, Collette Matthewman, Miriam Kent, Jenni Rant and Anne Osbourn
Plants, People, Planet (2020) 2: 602-613

“Postnatural Botany” rules booklet, plant discovery cards, and role-playing cars for “Explorer”, “Regulator” and “Artist”. Photo: Karen Ingram
Guest post by Karen Ingram, Creative Director
This past July, participants of the 2018 OpenPlant Forum went on an expedition where they discovered several new species of plants. Ok- they didn’t REALLY discover the plants, but they played a game that enacted the discovery of 18 new plants, from alpine to outback, at the conference dinner held at the Sainsbury Centre for Visual Arts (SVCA). The game, “Postnatural Botany” was inspired by Medieval Bestiaries, and the notion that explorers would describe the wildlife from their travels to people who had never seen such wildlife, as a means to share with the community.

Image courtesy of Rutgers University Honors College Instagram and Julia Buntaine.
“Postnatural Botany” has its origin as a workshop I created for 23 students in the Rutgers University Honors College and Douglass Residential College for women. The workshop took place in early 2018 and was part of a course, “Science/Art/Technology in New/York/City" taught by Julia Buntaine of the SciArt Center.
Originally dubbed “Postnatural Bestiary” and depicting a wide array of animals, I worked with Dr. Nicola Patron from the Earlham Institute to tailor the game to be plant focused, specifically for the Open Plant Forum.
The 2018 Open Plant Forum was hosted in Norwich, at the John Innes Centre. Norwich–which enjoyed great prosperity in the Middle ages–was the perfect backdrop for a game based on medieval bestiaries.
Each person was assigned a role as an “explorer”, “artist”, or “regulator” and worked in teams to produce artworks of each plant according to the rules of play. Patron selected a wide variety of plant life; Venus Flytrap, Rainbow Eucalyptus, Welwitschia, Jack-in-the-Pulpit, and Java Moss were all “discovered”, described with limited terminology and depicted with magic markers and imaginative minds.

Karen Ingram introducing the game to attendees of the Open Plant Forum. Photo: Nicola Patron
The “explorer” in each group was given an envelope that contained a card with the plant they had “discovered.” The card displayed a limited amount of information, including an image of the plant, the name, habitat, size, characteristics of its flower (if there was one), and information about a specific “outstanding feature” for each plant.
The “explorer” had to describe the plant they had discovered to an”artist” for visual interpretation. A “regulator” was also part of the game play, in order to ensure all parties followed the rules, of which there were many! Neither the artist nor the regulator were allowed to guess what the plant might be. The artist was not allowed to talk at all, primarily to keep them from guessing what the plant was.

Explorer Jenny Molloy gesticulates as she describes a Baobab tree to her team’s artists, Joanne Kamens and Dave Rejeski. Photo: Karen Ingram

Clockwise from bottom left: Francoise Kepes, Claudia Vickers, Steve Evans, Richard Sever, Anne Osbourn, Jim Haseloff, Peter Murray-Rust, Christina Smolke. Photo: Karen Ingram
I created a fictitious human-centric “public” that has no access to robust image catalogues and information we have via the internet, and no knowledge of the sciences. Because of these limitations, “explorers” had to use simplified terminology; leaning on familiar household objects and tools used by humans (purses, for example) and referencing the human body as a measurement unit. Professor George Lomonosoff from the John Innes Center remarked that it was a joy to describe a Rainbow Eucalyptus tree to his team’s “artists” as being “...as tall as twenty men!”

Jim Haseloff puts the final touches on his drawing. Photo: Nicola Patron
Simple explanations, gestures, analogies referencing common household objects, an abbreviated list of plant traits (stems, leaves, flowers, roots), as well as a few select domesticated plants were used in favor of scientific terminology.
The key objectives of the game were not so much to depict the plant correctly, but to work as a team: to communicate carefully on the part of the explorers, and to listen and interpret on the part of artists and regulators, and for everyone to have fun with plants.

An amazingly accurate rendering of a Sturt’s Desert Pea by Jan Lyczakowski, who exclaimed “I’ve never seen this plant before, but apparently I did a pretty well!” Photo: Nicola Patron
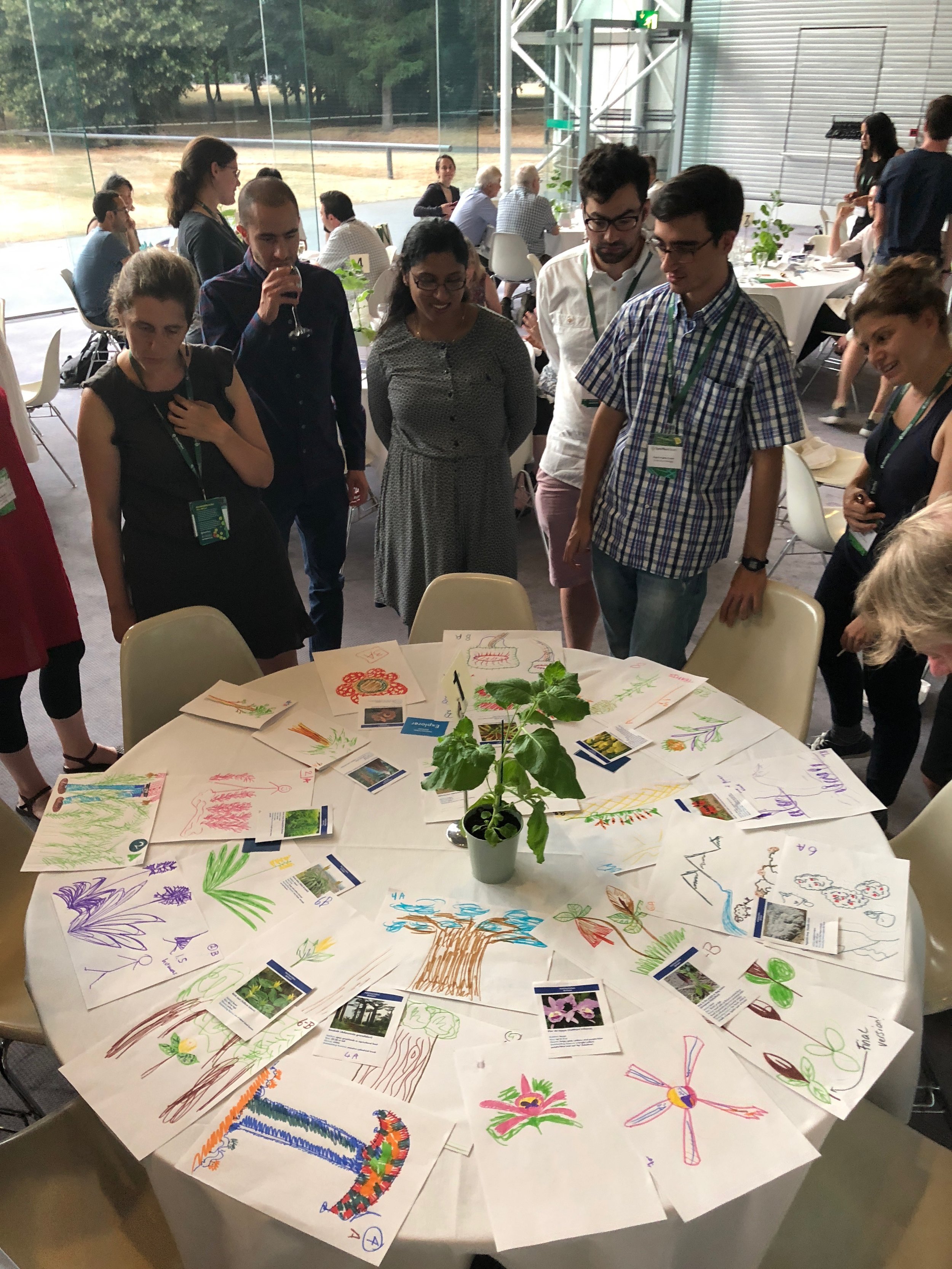

Open Plant Forum attendees marvel over a gallery of colorful creations. Photo: Nicola Patron
Colette Matthewman–whose winning drawings of a Jack-in-the-pulpit underwent several iterations before she was satisfied–shared her thoughts: “The OpenPlant Forum attracts a mutidisciplinary crowd, and this was a great game for breaking down barriers of language as we were all restricted to using very every-day words together with gestures to describe and understand the look of the plant – and the plants chosen looked really fantastical! As ‘an artist’ it was fascinating to see how I started to relate the explorer’s description to plants that I knew. This helped me to draw a reasonable likeness, but also limited my ability to take on board specific instructions from the explorer as they didn’t match the image in my head.”
The term “Postnatural” is defined as any organism altered by humans via selective breeding or genetic engineering. In the fable of this game, the plants and organisms are newly “discovered” by humans. Through the ages, plant collectors took their findings to new places for breeding and growth in new environments, altering the genetics and epigenetics of the plants forever. This calls to question, at what point of human intervention do organisms become “postnatural”? Once an organism is known and it is integrated into our lexicon; in a Bestiary as it was in the Middle Ages, domesticated to produce products for humans, or its genome sequenced, it is part of our human narrative. Fewer and fewer botanists get to experience the thrill of discovering a new plant species. And yet, through the discoveries of modern biology, humans are experiencing a new kinship with other organisms as we learn more about common biological processes and origins of life on earth. The gameplay of “Postnatural Botany” relies on observation, communication, listening, and interpretation; tools that we can all use to examine the potential impact of this kinship.
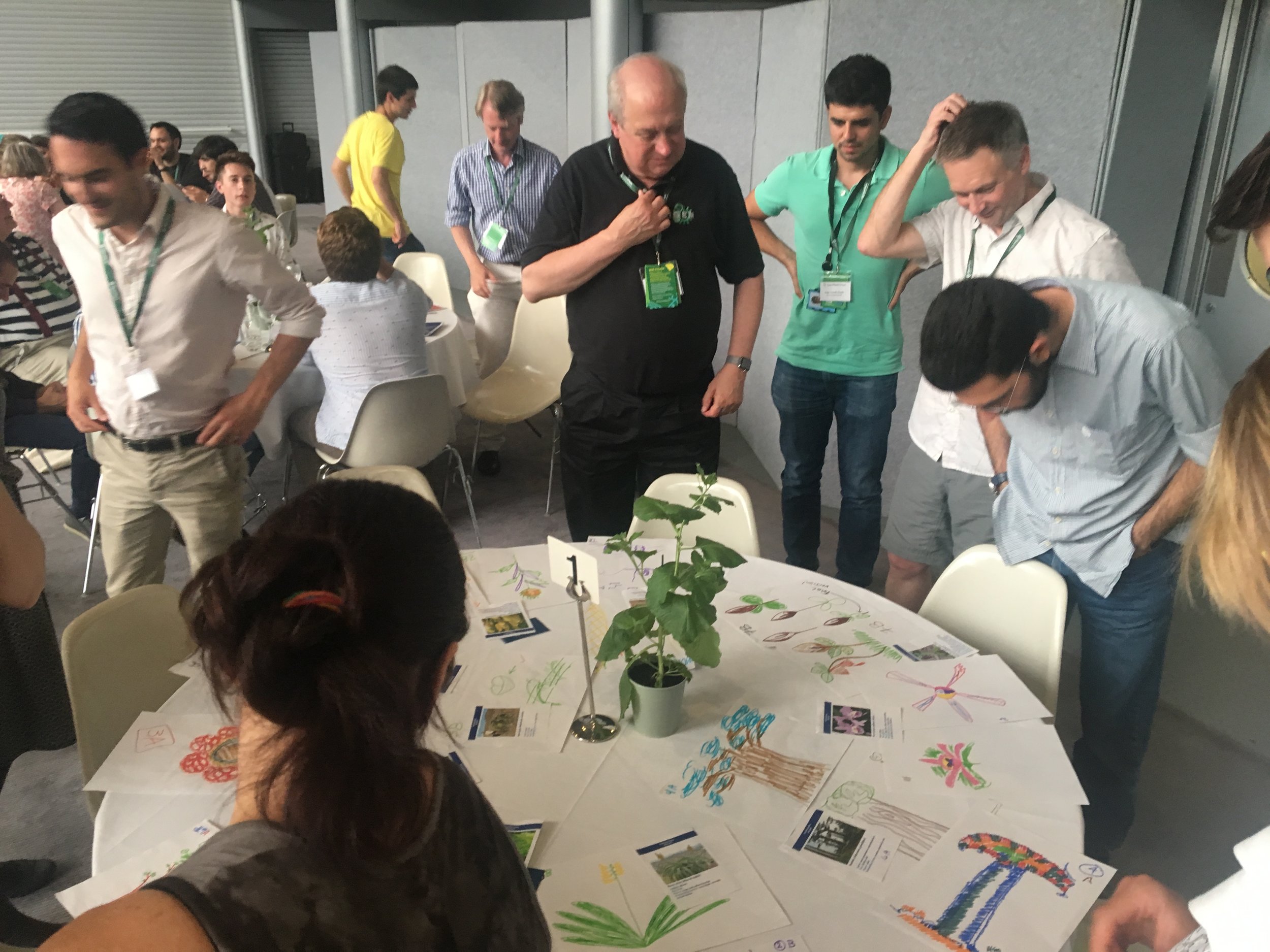

Markers down. Randy Rettberg, Roger Castells, and Ian Small survey the final pieces as people finish their pieces. Photo: Karen Ingram





Sketches show depictions of Venus Flytrap, Corpse flower, Java moss, Jack-in-the-pulpit (the winner), and Vegetable Sheep. Photos: Nicola Patron
Opening options for material transfer
Linda Kahl, Jennifer Molloy, Nicola Patron, Colette Matthewman, Jim Haseloff, David Grewal, Richard Johnson & Drew Endy
Nature Biotechnology 36:923–927 (2018)
In March 2017, Dr Colette Matthewman joined participants from industry, civil society, funding bodies and academia for an Industrial Dialogue on Synthetic Biology as part of the EU-funded SMART.map project (http://projectsmartmap.eu). The workshop was held at the Manchester Institute of Biotechnology.
The SMART.map project is developing RoadMAPs to Societal Mobilisation for the Advancement of Responsible Industrial Technologies. Its goal is to define and implement concrete roadmaps for the responsible development of technologies and services in three key time-changing fields: precision medicine, 3D printing in biomedicine and synthetic biology.
The Manchester dialogue provided a framework for discussions on the challenges facing synthetic biology companies, especially in the area of responsible research and innovation (RRI) and the development of concepts for tools that could help industry to engage with and integrate RRI into their synthetic biology working practice more easily. Read more in this blog and the video below.

OpenPlantand The Synthetic Biology SRI, Public Policy SRI and Faculty of Law co-organised a workshop held on 28 January 2016 on the openness of large bioresources in synthetic biology and genomics. The resulting report by Dr John Liddicoat and Dr Kathy Liddell has now been published on SSRN.
Research in synthetic biology and genomics depends on the use of collections of tissue and data, commonly known as bioresources. Substantial amounts of time and money are being spent on creating these bioresources and it is likely that significant scientific breakthroughs and development of end-products may be missed or delayed if the tissue and data in these resources are not shared. Accordingly, the ‘openness’ of these bioresources — in other words, the ability for other researchers to access, use, and share these resources (which is typically recorded in a bioresource’s IP and access policy) — is a key issue for the success of bioresource initiatives and the progress of synthetic biology and genomics.
There are, however, many different approaches to openness, and the development and dissemination of new knowledge are not necessarily advanced by distributing material at low cost or without any restrictions; time-limited rights of control (e.g. IP rights) may provide a useful incentive. It is a significant challenge to develop a fit-for-purpose openness policy that balances the advantages (and disadvantages) of different approaches to openness. The Workshop addressed this challenge by: reviewing openness policies adopted by large bioresources; eliciting ideas about access and intellectual property; debating the applicability of different openness policies; and identifying relevant areas for future research.
The report can be accessed here, and thanks and acknowledgments go to the Welcome ISSF and OpenPlant Fund. Both the Synthetic Biology SRI and OpenPlant were involved with co-organisation of funding along with Public Policy SRI and LML.
For more information please click here.
Image credit: Holly Gramazio via Flickr, licensed under CC-BY-SA 2.0
via Flickr, licensed under CC-BY-SA 2.0
Guest blog by PhD student Hannah Griffiths (John Innes Centre)

Have you heard of the Nagoya Protocol? Do you understand how and when Nagoya may affect your research? And most importantly do you know why Nagoya exists?
Let me put that another way...
Does your research involve working on any sort of biological material? Do you know where the material originated from? And do you have evidence that you are allowed to use it in your research?
Not sure? Then read on...
The Nagoya Protocol is a new international Access and Benefit Sharing (ABS) legislation, created to ensure that genetic resources are sourced and utilised fairly for providers and users.
Many people are not yet aware of Nagoya and, amongst those who are, there remains confusion regarding how and when it might be relevant to their research. In fact many non-commercial researchers assume that academic research will be exempt from this type of ABS legislation (which is not the case).
Nagoya is sometimes regarded warily as understanding whether your research falls in scope of Nagoya, and how to comply if it does, can be a daunting and time-consuming task. However, after attending the first regional workshop on Nagoya in the UK, myself and representatives from a variety of industries left understanding that fair ABS should be an integral part of research and that the Nagoya Protocol sets out bold plans to achieve this whilst simultaneously protecting biodiversity. The link between biodiversity and ABS may not be obvious, however it could be a crucial tool to protect biodiversity at a local level.

“…how easy it is to lose our connection with the natural world. Yet it is on this connection that the future of both humanity and the natural world will depend”
- David Attenborough (2016)
The presence of biodiversity in ecosystems, individuals and everything in between is critical for current and future life on earth. Natural and agricultural systems are more resilient to change when more biodiverse, and the unfathomable diversity of genetic elements in nature will be an essential resource for synthetic biology to innovate solutions to the World’s challenges. Yet, biodiversity has long suffered with intensive human population growth and urbanisation.
The 1992 Convention on Biological Diversity is a United Nations international treaty to address this and protect biodiversity. The Nagoya Protocol is a legal framework which entered into force over 20 years later, in 2014, to achieve the third and final objective of the convention;
“The fair and equitable sharing of the benefits arising out of the utilization of genetic resources”.
- Convention on Biological Diversity (1992)
The Nagoya Protocol describes how countries can exercise sovereign rights over genetic resources to ensure that providers of resources (or associated knowledge) receive a share of any benefit arising from their utilisation.
For the providers, the benefits received and their involvement in negotiating access demonstrates the high-value of their resources, which acts as a direct incentive to locally preserve such resources and consequently biodiversity.
Simultaneously, countries that become party to Nagoya agree to allow the utilisation of genetic resources under reasonable terms, which should ensure that the world’s genetic resources are actually available and accessible for research.
The UK became a party to the Nagoya protocol on the 22nd May 2016, the international day of biodiversity. Regulatory Delivery, the UK organisation that will be enforcing Nagoya, are currently raising awareness and understanding in relevant industries with activities such as this cross-sector workshop.
The workshop was very informative, with the Nagoya Protocol being thoroughly explained from the very basics to how to find answers to seemingly unanswerable questions, such as how to use the online chat feature on the Access and Benefit Sharing Clearing House (ABSCH).
The clarity of information helped remedy some common misconceptions. For instance, some people did not understand the incentive for the UK and EU to become party to Nagoya, as the association with biodiversity and ethics of fair sharing are not always immediately clear.
Further, there was a false impression that all benefits must be monetary, when in fact there is great flexibility and benefits are encouraged to be non-monetary, such as training and technology transfer. The main crux behind the “fair and equitable sharing” isn’t a monetary value but something that is negotiated and agreed fairly between the user and provider before sharing takes place. For many researchers this may already occur in an informal way, the Nagoya protocol just formalises the process and provides legal certainty.
Speakers from various industries also shared their views on Nagoya. For some of the speakers from huge historic collections, such as Natural History Museum and Kew Gardens, Nagoya implementation meant reassessing and tightening ABS systems already in place, which was seen as a positive action.
A speaker from AstraZeneca shared the systems they had put in place throughout the company to implement and raise awareness of Nagoya, including a great promotional video.

ABSCH opening webpage https://absch.cbd.int/
The workshop also raised a number of interesting discussion points about the challenges of Nagoya. For instance; the enormity of the task of ensuring all future research is Nagoya compliant, the difficulties for research that bridges the gap between non-commercial and commercial research, and that small pilot studies may not have the resources to implement Nagoya.
The relevant UK and EU authorities are aware of the challenges of Nagoya, and eager for feedback and suggestions of best practices to incorporate into future legislation and Nagoya implementation.
Therefore, for most of the challenges discussed positive ideas and solutions could be suggested. Some ideas raised at the workshop included; scientific journals requiring evidence of Nagoya competence before accepting papers, funding bodies providing small amounts of money to allow pilot studies to be Nagoya competent, and the idea of Nagoya competent registered collections of genetic resources (which is already an article in EU legislation).
It will take time for implementing Nagoya to become a clear and easy process, however, for the principles of undertaking fair research and moreover protecting biodiversity, it will be crucial and undoubtedly worth the effort.
This article is authored by Gerant Parry and originally appeared on the GARNet blog, it is republished with permission.

Photo credit: the iGEM Foundation and Justin Knight
Take a look over there! As even the name suggests, the iGEM Giant Jamboree is a conference like no other.
Consider that there are 2500 mostly undergraduate students from all around the world, the vast majority of them at their first conference and each giving presentations that are being critically assessed. This provides a clue as to the kind of frenetic and excited energy that characterises this event.
For those a little confused, the International Genetically Engineered Machine Foundation oversees and organizes iGEM, which is synthetic biology competition for groups of participants who are usually hosted by academic institutions. The basic idea is that a group of students works through the summer on a completely novel project that conforms to the principles of synthetic biology, before presenting it in the aforementioned Giant Jamboree.
As this is a competition, each project is judged on metrics that assess many aspects of the teams work. These include the contribution of biobricks to the iGEM registry (an impressive selection of molecular parts that are held within a standardised plasmid), the development of their novel project, initiating collaborations with other teams and their attempts to integrate human practices and public engagement into their project. By meeting certain criteria each team is eligible for Gold, Silver or Bronze medals alongside special prizes for different project categories.
Given registrations, student stipends, research expenses, travel and accommodation, putting forward even a small team can stretch to at least £20K. Therefore this is not a something to be taken lightly. To this financial requirement must be added the time donated by a team of instructors and advisors that support the students. However regardless of the cost, one thing is certain; for those students who participate, attend, present and are inspired by the Jamboree, it can be a career-defining moment.
Plant Synthetic Biology can make for a challenging summer!
Plant experimental chassis have not been widely used during the ten years of the iGEM competition where bacteria, yeast, mammalian cell lines or cell-free systems offer time efficient alternatives for the usual 10-week research period. However the iGEM foundation, alongside a group of committed advocates have recently developed the Phytobricks cloning standard, which is based on a recently published standard syntax within the Golden Gate cloning system. The aim is to lower the barrier of accessibility for teams to start plant projects and the evidence from this years competition seems to suggest that this is slowly happening. The 2016 iGEM team from Valencia-UPV is advised by plant synthetic biologist Diego Orzaez and their project submitted phytobricks for the expression of a split Cas9 system. They showed that the two halves of the Cas9 protein could reconstitute and was active in a tobacco expression system. They have documented this work on their Parts pages and this is hopefully a resource that will be used by future iGEM teams. Their team was very successful at the jamboree, winning a gold medal alongside specific awards for the best hardware and software.
Another successful team with a plant project was from SCAU-China who had, over the course of at least two years, added an additional two genes to conventional Golden rice. This produces a ‘brown rice’ that produces the natural keto-carotenoid Astaxanthin, which is thought to have beneficial anti-oxidant properties. This is clearly a significant research project that has been badged with the iGEM logo and as such was very positively received by the judges. Although they did not submit parts in the Phytobricks standard it was exciting to see such a potentially high profile plant-project feature at the jamboree. These projects are well deserving of their awards and their work builds upon years of expertise contained within the supporting labs. This highlights one of the challenges for the competitive element of iGEM; namely how teams can be equally judged when they have hugely varying levels of support. Fortunately it appears that this is not a significant issue as each team is able to take positives from their own performances and are happy to celebrate the excellent projects that they each had individually put together.
Remarkably the iGEM competition includes at least 30 high school teams and one of these, GDSYZX in China, worked with plant light responsive promoters that they added to the Parts Registry.
Algae on the rise.
A number of teams including Cambridge-JIC, Linkoping University in Sweden and USP_UNIFESP in Brazil used the algae Chlamydomonas_reinhardtii in their projects. Cambridge team had most success in their project that generated a set of parts in the Phytobrick standard that can be used in future algal projects. In addition they created a remarkable blueprint for the production of a prototype Genegun for plant transformation, costing just £300, making it accessible for less well funded labs. The other two teams mentioned above were hoping to use Chlamydomonas to produce either biofuels or spider silk protein and although the ambition of both projects outstripped their achievements this year, iGEM is all about thinking big: sometimes it works, sometimes not!
The team from Pretoria in South Africa took on an extremely ambitious plan called WattsApatmer to create ‘plant batteries’ by using short aptamers to attach either photosystem II or a laccase enzyme to either pole of an electrical circuit held within a novel graphene scaffold. The students made some progress with this and the project serves to highlight the blue-sky thinking that undergraduate students undertake as part of this competition. It is clearly difficult to make enormous progress over a summer project but there were so many amazing project ideas on display at this iGEM I hope that the host institutions can find finances to develop some of these ideas so that some can come to fruition to add value to the time already committed to these projects.
Europe on Top
From a UK and European perspective the iGEM jamboree was a huge success with Imperial College and LMU TU Munich taking the overall undergrad and overgrad awards respectively, with remarkable projects that highlighted the talent of their students and the level of support their receive from their host institutions.
The UK was represented by over 20 teams, the third most numerous behind the USA and China. Aside from Imperial College, the teams from Exeter, Dundee, Dundee Schools, Cambridge-JIC, Oxford, Sheffield, UCL, Glasgow and Manchester gained Gold medals. There is little doubt that the UK is developing a cohort of talented synthetic biologists who will be the research leaders of the future.
Overall we look forward to seeing the number of plant projects increase over the years to come. The development of the Phytobrick standard will undoubtedly help in this goal for students to come up with ideas to test the possibility of using plants in their projects.
There are exciting times ahead for plant synthetic biology!
Betty and Gordon Moore Library Wilberforce Road Cambridge CB3 oWD
OpenCon 2016 is the student and early career academic professional conference on Open Access, Open Education, and Open Data being held in Washington, DC.
The OpenCon 2016 Cambridge satellite event will bring together students, early career academic professionals and open advocates from around Cambridge (although anyone from the surrounding areas are welcome to join us!)
This year’s theme is Building Impact Through Openness.
Our goal is to support and build the open community in Cambridge. We want to empower attendees to make a difference in their respective fields through open research, data, education and access.
This year’s committee have brought together a sensational program of world leading speakers but there is also time scheduled for focus group discussion around actions that we can take to make a change in the world.
Furthermore there’s ample time set aside in the day to ensure all the attendees will be able to share their ongoing activities (in the “silent” unconference) and hopefully build new collaborations for the future.
You can follow the main OpenCon event in held in Washington DC #OpenCon, and you can join the discussion around the Cambridge satellite event at #OpenConCam2016.
We look forward to seeing you there! Please reach out if you have any questions.
For more information and booking, click here.
The event is organised by Kirstie Whitaker, on behalf of the OpenCon Cambridge organising committee
Monday 17 October 2016, 18:00Panton Arms 43 Panton Street CB2 1HL, Cambridge.
Café Synthetique is the monthly meetup for the Cambridge synthetic biology community with informal talks, discussion and pub snacks.
iGEM Cambridge team 2016 are taking part in the prestigious iGEM synthetic biology competition. Focusing on chloroplast engineering, the team are interested in using chloroplasts to produce metabolic products, such as biofuels and edible vaccines.
Michelle L. Oyen is a Reader in the Bioengineering in the Mechanics and Materials Division and the Bioengineering research group in the Cambridge University Engineering Department. Her work involves research on mechanical behavior in biological materials with many of her projects having a distinct biomedical focus.
"InstaCHLAM, a toolbox for chloroplast engineering"
iGEM team 2016, Cambridge, Department of Plant Sciences, University of Cambridge
"Biomimetic materials for structural applications"
Michelle L. Oyen, Reader, Cambridge University Engineering Dept., The Nanoscience Centre
Please join us in the Panton Arms pub at 6pm.
More information about this event…
Image credit: Photo from BASF on Flicker, licensed under cc by-nc-nd/2.0/-Image source
18 October 2016, 7:30pm - 9pm followed by drinks reception
Dr Kevin Esvelt (MIT Media Lab) and Professor Luke Alphey (Pirbright Institute, founder of Oxitec Ltd) examine the science, ethics and regulation of genetic engineering to control mosquito-borne disease. What promise does this emerging technology hold and how do we ensure it is used responsibly?
Biologists can now design genetic systems that engineer evolution in powerful ways with social, legal, ethical and environmental implications for our future. Mosquito populations can already be engineered using cutting edge techniques to drastically reduce their numbers or make them resistant to transmitting diseases like malaria, dengue or the emerging Zika virus.
Synthetic biologist Dr Kevin Esvelt (MIT Media Lab) will introduce his work on gene drive systems which rapidly spread malaria resistance within populations while Professor Luke Alphey (Pirbright Institute) will discuss his work founding Oxitec, a UK company that was the first to release genetically modified male mosquitoes whose offspring fail to reproduce, leading to dramatic reductions in numbers.
What safeguards and regulations are required to ensure responsible use of such technologies? What does it mean for humans to use nature's tools in this way? How do we balance the direct benefits for global health with any risks to our shared environment?
Talks and dialogue on the idea of sculpting evolution will be followed by a drinks reception.

Tuesday, November 8, 2016, 6:30 PMto8:00 PM
Old Divinity School, St John's College, St Johns St, Cambridge CB2 1TP, Cambridge
Professor Ron Weiss (MIT) introduces the design and implementation of synthetic gene circuits in mammalian systems, exploring the potential of this approach in regenerative medicine and stem cell engineering. The talk and dialogue will be followed by a wine reception and delicious finger buffet.
Professor Ron Weiss (MIT) is a pioneer of synthetic biology and is currently Professor of Biological Engineering at MIT in the Department of Biological Engineering and the Department of Electrical Engineering and Computer Science.
The Weiss lab uses computer engineering principles of abstraction, composition, and interface specifications to program cells with sensors and actuators precisely controlled by analog and digital logic circuitry encoded in synthetic gene networks. These circuits can be used to control the behaviour of individual and aggregated cells and from early work in bacteria, the lab has more recently explored transcriptional regulation in mammalian cells. Professor Weiss’s research has traced a journey from genetic parts to modules and is now devising therapeutic systems that more reliably direct stem cells to create new tissues. This work aims to move towards replacing the cells lost to disease or injury, pushing the frontiers of the nascent field of synthetic morphogenesis. In this talk, we will explore the potential of synthetic biology as an approach in regenerative medicine and stem cell engineering.
The talk and dialogue will be followed by a wine reception and delicious finger buffet.
Registration: £10/£5
This event is organised by the Synthetic Biology Strategic Research Initiative as part of our Michaelmas Term 2016 SynBio Forum. For more events please visit:
Sculpting evolution: engineering biology to address global disease challenges Venue: Howard Building, Downing College, Cambridge
Date: 18 October 2016, 7:30pm - 9pm followed by drinks reception
Register: http://tiny.cc/synbioforum-18Oct2016 Dr Kevin Esvelt (MIT Media Lab) and Professor Luke Alphey (Pirbright Insitute, founder of Oxitec Ltd) examine the science, ethics and regulation of genetic engineering to control mosquito-borne disease. What promise does this emerging technology hold and how do we ensure it is used responsibly?
Programmable biology in the test tube
Venue: Department of Plant Sciences, Downing Site
Date: 19 October 2016, 09:00-17:00, including talks and practical
Register: http://tiny.cc/synbioforum-19Oct2016 Synthetic gene circuits can be used to generate rapid and low-cost paper-based diagnostics for diseases including Zika and Ebola. Dr Vincent Noireaux (University of Minnesota), Dr Nick Rollins (Cambridge Consultants) and Dr Fernan Federici (Pontificia Universidad Católica de Chile and University of Cambridge) present the technology and its disruptive implications during these lunchtime seminars and a hands-on prototyping workshop (application required). The OpenPlant Fund will launch a linked call for mini-grants to support interdisciplinary collaborations on the theme of in vitro synthetic biology.
Synthetic biology for regenerative medicine
Venue: Old Divinity School, St John’s College, St Johns St, Cambridge CB2 1TP
Date: 8 Nov 2016, 18:30 - 20:00 followed by networking reception with buffet
Registration (£10/£5): Link to be posted to http://www.synbio.cam.ac.uk when live Professor Ron Weiss (MIT) introduces the design and implementation of synthetic gene circuits in mammalian systems, exploring the potential of this approach in regenerative medicine and stem cell engineering. The talk and dialogue will be followed by a wine reception and delicious finger buffet.

Have you ever wondered how new ideas can help people in the developing world? Are you interested in what Cambridge can do to help? If so, the Centre for Global Equality Development i-Teams programme is for you!The University of Cambridge SynBio SRI has put forward a synthetic biology-based project for the Development i-Teams Michaelmas 2016 'Exploring potential global health applications of cell-free extracts for rapid, low-cost, paper-based diagnostics', mentored by OpenPlant Fellow Dr. Fernan Federici. Find more information and how to apply below!
Researcher : Dr. Fernan Federici, Plant Sciences, University of Cambridge and Pontificia Universidad Catolica de Chile
In vitro synthetic biology uses cell-free extracts from bacteria or other organisms to which DNA sequences encoding genetic circuits with useful functions are added and expressed. For example, a molecular sensor for Zika viral RNA or a pollutant heavy metal could be designed to produce a signal that regulates output of a measurable response, like high levels of a coloured chromoprotein. Beyond this simple example, DNA-encoded ‘logic gates’ could be constructed that respond in different ways to particular combinations of inputs or provide quantitative results.
Recent work combining this emerging technology with paper-based microfluidics has delivered rapid, low-cost paper tests for Ebola and Zika Virus and small molecule sensors such as glucose assays, which are stable in dried form for at least one year. As no genetic modification is involved, and a fully equipped lab is not required once the cell-free extract is produced and stably dried down, this technology is far more accessible to researchers in low-resource settings than in vivo synthetic biology. The initial cost of engineering biosensors is lowered very significantly.
The potential applications of this technology are directly relevant to diagnosing health and environmental problems faced by people developing countries. Many such problems pose significant challenges to their welfare and economic development e.g. pollution, tropical infectious diseases, animal disease, soil health. By promoting the development of a low-cost, low-resource technology platform the intention is to build capacity in-country for prototyping solutions to challenges identified as priorities locally.
The inventor of the technology, Dr. Fernan Federici, is working on open technologies including hardware and DNA parts that would further increase the ease and suitability of these cell-free systems for research, development and applied use in the global South. With reduced IP encumbrance, it is hoped that knowledge transfer can be accelerated and barriers to access reduced.
The Development i-Team will need to investigate two separate questions relevant to this technology.
First they will investigate the likely applications of paper-based synthetic gene networks in the developing world, and in particular will need to identify areas where local problems are not addressed by existing solutions and there is scope for developing local capacity for research in this area.
Secondly they will explore the commercial implications of building diagnostics based on open technologies that are freely shared and not protected by patent. How does this make a difference in the global South? How does it alter the typical value chain for such technologies? Could open approaches confer benefits in terms of access for the bottom three billion and in what contexts? In particular, are there any mechanisms of sharing IP with the local community which is directly affected by the problem?
The Centre for Global Equality and i-Teams are running the "Development i-Teams" programme for the fourth time this Michaelmas term. Teams will investigate ways in which real Cambridge innovations could be used in the developing world to improve people’s lives in a sustainable way.
This term’s projects are:
Development i-Teams is open to all students (undergraduates and post-graduates), post-docs and staff, as well as all members of the Centre for Global Equality - anyone with an interest in how technology can make the world a better place for the world's poorest.
The course runs on Tuesday evenings from the 18th of October to the 22nd of November, and there will be approximately 4 hours of individual work needed each week, mostly involving gathering real-world feedback from experts in international development.
For more details and to apply for a place on a team, see http://iteamsonline.org
For more information about the work of the Centre for Global Equality see http://centreforglobalequality.org

The Nuffield Council on Bioethics has today published the first findings of its programme of work looking at the recent and potential impact of recent advances in genome editing such as the CRISPR-Cas9 system across many areas of biological research.
The Council found evidence that, given its technical advantages and rates of uptake, genome editing is already having an almost unprecedented impact in research. The Council considered factors such as the extent to which the ethical questions raised by applications of the technology are novel, the likelihood of imminent advances in these areas and the possible effects of these advances in fields such as health care, food production, industry and public health.
Genome editing techniques are an essential tool for synthetic biology and while centred around more standard forms of single gene editing, the report acknowledges the field (sections 7.3-7.6) and the use of gene editing in food crops (5.1-5.17)
"Synthetic biologists are self-consciously elaborating a novel field. They see the field as transforming biology as a practical discipline, not only in relation to the adoption of technical innovations, but also epistemically and institutionally (breaking down disciplinary barriers and reimagining biology as an engineering discipline), and socially and politically (e.g. the desire to build a community and to inculcate certain norms, including those of open source publication and responsible innovation practices). While, undoubtedly, genome editing has given a fillip to synthetic biology it does not, however, seem to have the same rhetorical significance here as in other areas of biology. This might be partly attributable to the fact that the natural reservoir of metaphor for synthetic biology is technical (engineering, construction) rather than textual (editing).
Synthetic biology does, however, offer an insight into possible ways of approaching genome editing as an innovation within research and industry that is essentially different to the translational approaches of biomedicine or, again, public health innovations. Owing, in part, to the different cultures that are integral to synthetic biology (e.g. that of computer science) and in part to lessons about innovation learned from the observation of other fields (e.g. nanotechnology), it has been common for synthetic biologists to adopt responsible innovation practices from the outset. These tend to see ethical reflection and social engagement as longitudinally integral to their practice (‘ethical by design’), as both guiding and governing research, rather than as challenges or decisions to be addressed at particular stages."
"Genome editing is currently used in research into plant breeding. Possible commercial uses include improvements in yield and pest resistance, increased drought tolerance, and increased nutritional benefit.
The impact of genome editing techniques is perhaps less revolutionary in plants than in humans, given the already long history of breeding strategies that have changed the genetic characteristics of virtually all crops – including selective breeding and first generation ‘genetically modified’ plants (mainly involving the insertion of genes that do not naturally occur in those plants).
However, genome editing could significantly speed up the progress of breeding programmes. It is thought that genome editing could reduce the time needed to generate the desired genetic characteristics in a plant population from 7-25 years to as few as 2-3 years since its target specificity effectively bypasses the need to go through a number of plant generations to achieve a particular genetic combination.
Depending on the regulatory and economic conditions, it could open up the field to smaller companies and, potentially, drive the development of characteristics other than the main commercially important traits like herbicide resistance."

The Department of Plant Sciences hosts significant synthetic biology research including OpenPlant and the SynBio SRI. It is currently welcoming independent research fellows who wish to apply to join the Department.
We welcome approaches to host candidates who are applying for independent Fellowships, such as Royal Society University Research Fellowships, BBSRC Sir David Phillips, NERC or EU Marie Curie awards. The Department has an excellent tradition for supporting Research Fellows in terms of providing financial support for laboratories and equipment, and PhD studentship opportunities, as well as promoting career development and collaborative expertise.
Potential candidates should contact the Head of Department, Professor Alison Smith (hod@plantsci.cam.ac.uk) or Department Administrator, Catherine Butler (cek31@cam.ac.uk) initially, after which they should approach Group Leaders from the Department (http://www.plantsci.cam.ac.uk/research) who are most closely allied to their area of research interest. Applications will be co-ordinated by the Department Resources Committee. Contact should be made at least 6 months before any deadline for submission. We welcome applications from individuals who wish to be considered for part-time or other flexible working arrangements.
The Department of Plant Sciences sits within the School of Biological Sciences, with undergraduate teaching integrated through the Natural Sciences Tripos (NST). Postgraduate recruitment and training is offered either directly or indirectly via BBSRC or NERC Doctoral Training Programmes (http://www.plantsci.cam.ac.uk/grads/). The Department maintains teaching and research specialisms across a wide range of plant science disciplines (from molecular and developmental biology, through cell-signalling, biochemistry and physiology to epidemiology, ecology and ecosystem modelling) with 18 academic staff leading active research groups, 4 independent research fellows (funded by the Royal Society, EC and NERC), 4 senior research associates, 55 post-doctoral researchers and 42 support staff. Research grant income in 2014/15 was £6M, with the Department currently administering a total of 68 grants with a combined value of £26.5M from a variety of sources, including research councils, Royal Society, charities, EU, industry and government agencies.

Many early efforts of synthetic biology have focussed on the engineering of microbes, especially for the growing biotech industry. In contrast to single cell microbes, multi-cellular organisms such as plants present a higher level complexity, take longer to engineer, and the regulatory system can be a tough and time consuming to navigate - but there are huge opportunities for delivering social, environmental and economic benefits through efforts to reprogramme plants and agriculture. They come with their own distinct set of ethical, legal, social and economic questions. The above were topics central to discussions at the 2016 OpenPlant Forum. Over one hundred people from various disciplines assembled to hear about some of the recent advances in crop and feedstock engineering, discover the latest tools to support innovation in this field, and to reflect on and discuss the ethical, legal, social, and economic questions.
Events kicked off at the John Innes Conference Centre, Norwich, with a networking evening and industry showcase, including two exciting new local developments: Martin Stocks (Plant BioScience Ltd) talked about Leaf Systems®, a translational facility being built to scale up protein and chemical production in plants; and Tony West gave a preview of the new DNA Foundry at the Earlham Institute, which has since been officially launched.
The first full day of the Forum opened with a double bill of keynotes from Allan Green (CSIRO) and Jonathan Napier (Rothamsted) talking about their impressive efforts engineering oilseed crops. It continued with a case study of AB Sugar's Wissington sugarbeet processing site, providing an inspiring processing model for maximising production from a feedstock and it's byproducts. This was followed by a cross-discipline exploration of some recent advances and future opportunities for reprogramming agriculture. In the final session of the day, Spencer Adler (Bioeconomy Capital) gave an investors perspective, followed by a lively debate on the ethical, legal, social and economic considerations of developments in this area. Discussions continued into the night at the conference dinner.
Day two grounded the discussions back in the technical, with a focus on tools to support synthetic biology, especially in plants. The day started with Tom Knight opening the curtains to an exhilarating view of Ginkgo Bioworks and some of their latest developments. Moving back to plant chassis, advances establishing the liverwort Marchantia as a simple plant chassis were showcased alongside work developing tools and methods for other plant chassis. The final session of the event focussed on tools to enable innovation through sharing of knowledge, data and materials - a key focus of the OpenPlant Synthetic Biology Research Centre.
Steven Burgess and Cindy Chan have published a detailed write-up of the OpenPlant Forum on the PLOS Synbio Community blog: Seven Developments in SynBio: Science, Patents and Ethics | OpenPlant Forum 2016
Blog post written by Colette Matthewman Photos by Matt Heaton
For more information, see the link: http://www.stw.nl/nl/content/biotechnology-and-safety
A matchmaking event is included in the process and will take place on 15 September.
The OpenPlant-supported Cambridge-JIC iGEM Team are exploring open source synthetic biology tools for chloroplast engineering in algae. The following was authored by Cambridge JIC- iGEM team member Claire Restarick and is reposted with permission from the Cambridge Consultants blog. Since our initial blog post, we’ve spent many hours finalising designs for our genetic assemblies, which are now in the process of being synthesised. Once these are complete, we will begin the challenging task of completing four rounds of experiments before our deadline in September.
While our biologists are making significant headway in the lab, there have also been advancements on the hardware and engineering side. Our designs for low-cost, open-source lab equipment to support our Chlamydomonas transformation protocol have started to take shape – with the first stages of assembly taking place. This equipment will include a growth facility with light control, temperature regulation and imaging capabilities (linked to a dedicated Twitter account, @RPi_camigem2016), as well as a gene gun which, if successful, will transform cells by firing DNA-coated tungsten microparticles directly into them at high pressure.

The decision to make our hardware low cost and open source developed from recent trips to publicise our project at synthetic biology conferences in Paris and Norwich. At the Bio NightScience event hosted by the Centre de Recherches Interdisciplinaires (CRI) at the Cité des Sciences et de l’Industry, we presented our project to the conference and heard from many projects originating from Makespaces – collaborative community labs with little-to-no budget. We found its use of plant synthetic biology was hindered by the high cost of commercial equipment to culture and transform plant and algal cells. This inspired us to design low-cost equipment, which could make the area of plant synthetic biology more accessible to these creative workspaces, and other small research institutions.
The issue of documentation for open-source hardware for synthetic biology was raised repeatedly during the Open Plant Forum, hosted by the John Innes Centre in Norwich. A lack of clear, detailed protocols online makes it near impossible for the average novice builder to construct these devices. Having struggled ourselves to find appropriate parts and clear designs online, we have placed a focus on thoroughly documenting our designs, to make our open-source designs truly accessible for everyone.
To support both the hardware and biology aspects of the project, we have also continued our work on mathematical modelling, developing an open-source, integrated, kinetic model of Cas9-mediated gene insertion. We also held our first meeting with the director of the Cambridge-based Centre for Global Equality, to begin developing the human practices element of our project – understanding its impact and integrating this within the design and aims of the different parts.
Now past the halfway point of our project’s timeline, we feel well on track to meeting our project’s ambitious goals. Thanks to the continued support of our advisors at the Plant Sciences Department, and specialist advice from Cambridge Consultants, all aspects of our project are developing the potential to have an impact on both the scientific and non-scientific communities.
Note from Cambridge ConsultantsSynthetic biology has huge potential to solve many of today’s critical challenges in healthcare, agriculture, energy and the environment. That’s why Cambridge Consultants has decided to sponsor the Cambridge University team at iGEM 2016 – the international genetically engineered machine competition run by MIT. As part of our sponsorship, we are acting as mentors – giving the team access to more than 700 Cambridge Consultants engineers and scientists worldwide to help solve problems during this year’s project.
The iGEM team is also grateful for support from:
CamCreatives event - sign up here
Wednesday 29th June, 7-9pm Hot Numbers Cafe Dale's Brewery, Gwydir St, Cambridge, CB1 2LJ
Title: Reframe: Get unstuck and create breakthrough ideas
Einstein defined insanity as “doing the same thing and expecting different results” Simply changing your mindset and behaviour can create better outcomes for you and your team. Experience for yourself the world’s most successful creative leadership programme to solve real challenges here in Cambridge.
[spacer height="20px"] THNK School of Creative Leadership ’s mission is to catalyze innovative solutions to the world’s challenges through experiential learning programmes. At this workshop you will learn how you can adapt your own mindset to get the most out of team working. You’ll experience an interactive session using THNK methodology to help generate new perspectives and come up with new solutions. You’ll apply the methodology to a real challenge by a local group and there’ll be a prize for the winning solution. John Monks is a lifelong changemaker, startup cofounder and digital transformation expert. He completed the THNK Leadership Programme in 2015 and is on a mission to make the world a better connected place. John found himself drawn to purposeled work and is committed to bringing knowledge and tools into the world’s leading companies. Tonight John will present you with key challenges facing our city and introduce a key THNK methodology to apply to these challenges live. You will leave enabled to lead in a more creative way in your work and have greater impact. John will be joined by Nicky Shepard, founder of Cambridge Style Week, who is committed to building a creative space for Cambridge a community with the space, tools and resources to leverage the creativity in our City and have lasting impact. Poet and food activist Peter Bickerton will also be on hand to entertain us.
[spacer height="20px"] John is cofounder of DOTWORKS , a startup aiming to make the world a betterconnected place, and Trustee of ActionAid, and international charity working with the poorest women and children in the world, changing their lives for good. Nicky Shepard is a creative thinker and leader, the founder of Cambridge Style Week and committed to helping people build successful businesses. Her experience runs across drama and performing arts, events management, marketing and community building. Peter Bickerton is a science communicator, poet and writer based in Norwich. Responsible for promoting the research of The Genome Analysis Centre and the wider scientific community, he is also an enthusiastic ambassador for Thought for Food and a passionate insect eating connoisseur.

The first three calls for proposals to the Global Challenges Research Fund are now open. The Fund, involving BBSRC, MRC, AHRC, ESRC and NERC, was created to ensure that UK research takes a leading role in addressing the problems faced by developing countries. Foundation Awards of up to £600,000 are available for multidisciplinary research in three areas:
Deadline for outline applications: 4pm, 22 June 2016 Awards would start on 1 April 2017, for applications for proposals of ~24 months’ duration.
All research funded through these awards will be part of the UK’s Official Development Assistance (ODA). Applications must be primarily relevant to near-term or long-term benefits to the health or prosperity of Low or Middle Income Countries.
Note for Cambridge researchers:Cambridge Global Food Security and CambPlants coordinate a large network and can help you find the collaborators you need across the University for a multi-disciplinary proposal that meets the funding criteria. In anticipation of further funding calls in this area with a short turnaround (GCRF, Newton Fund, etc),the Global Food Security SRI are planning a series of Sandpit events on themes within Global Food Security to support Cambridge academics in generating and developing new ideas, finding collaborators (both within Cambridge and from external organisations), and submitting bids. Please let Jacqueline Garget – Coordinator of the Cambridge Global Food Security SRI - know if you would like to be involved in the events or if any research ideas you’re thinking of submitting that fall within the remit of the Initiative (www.globalfood.cam.ac.uk).